Use Case 1.1
Motivation
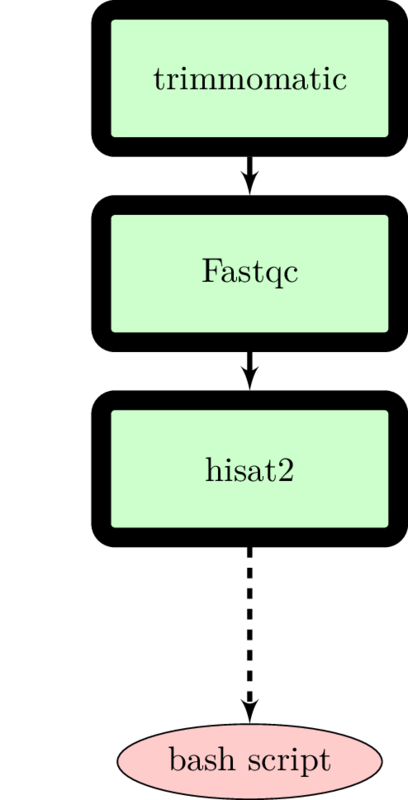
The Docker philosophy states that every container should serve a single function. This makes it easier to update a single tool without worrying how that affects the overall project. For instance, say you want to redo your analysis, but this time you want to use a different read mapper, say tophat2 instead of hisat2. Containerizing your individual tools allows both modification (grabbing a different Docker image) and portability between different operating systems.
Requirements
sudo docker pull quay.io/biocontainers/trimmomatic:0.36--5
sudo docker pull fjukstad/fastqc
sudo docker pull limesbonn/hisat2
Usage
Make sure that the input files are in your current working directory before running.
Usage: ./uc-1.1.sh -r1 <READ1.fastq.gz> -r2 <READ2.fastq.gz> -a <adapters.fa> -g <ref.fa>
Options:
-r1 --read1 a read1 gzipped FASTQ file
-r2 --read2 a read2 gzipped FASTQ file
-a --adapters an adapters FASTA file
-t --trimfile a trim file for trimmomatic
-g --genome-ref a reference genome FASTA file
Example:
./uc-1.1.sh -r1 sample_1_R1.fq.gz -r2 sample_1_R2.fq.gz -a Adapters.fa -g reference_genome.fa -t testfile_trimmomatic.txt
